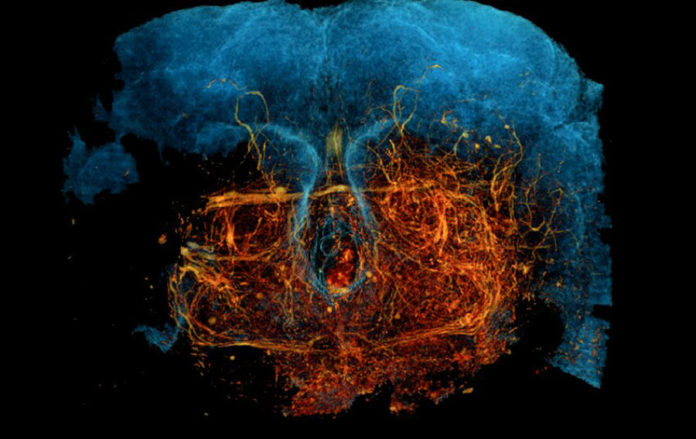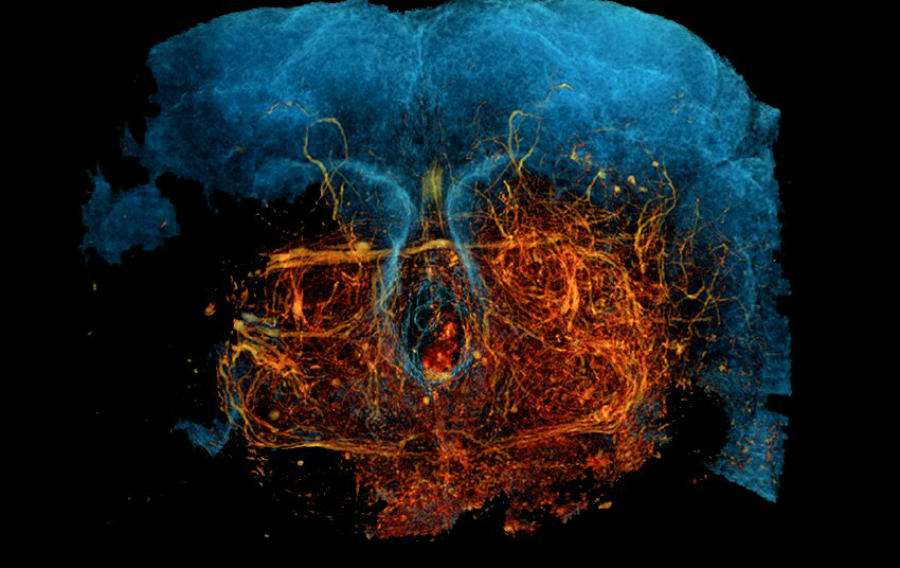
(Credit: Kuan et al, 2020)
Imaging the very complex web of nerves in the brain has been made possible by a team of scientists at Harvard Medical School. The new technique is called, “X-ray holographic nano-tomography (XNH)” and can map neural circuits in high-resolution imaging.
The human brain has over 100 billion neurons with over 100 trillion nerve connections. Having a precise map of the brain of an individual has long been the Holy grail of neuroscience. But if it could be precisely achieved, scientists could gain access to study our body functions, thoughts, memories or behaviours in unprecedented levels.
Reporting their latest discovery in the journal Nature, the team explains that their technique has so far able to successfully image neural pathways in a mouse brain and also in the fruit fly.
The technique involves artificial intelligence (AI) to construct 3D imaging in the brain which gives the opportunity to clearly distinguish individual neurons and their pathways leading to the muscle. And it is said that the new X-ray microscopic technique is far ahead than Electron microscopic (EM) techniques which are in use for such applications today. The new technique is able to image larger parts of nerve tissue in a shorter time.
The new technique in use
In previous attempts, scientists were able to fully map a fruit fly brain. This was done by “serial slices of the brain” which are a 1000 times thinner than a human hair strand. The slices were then imaged using EM. The separated images were then combined together for analysis. But techniques like these could be very costly and time-consuming which also requires expert knowledge.
But in the new technique XNH, it takes rotating X-ray images of the body/ tissue which is similar to a CT scan. But here it used high energy x-ray beams generated by the ESRF’s synchrotron, where electrons accelerate to the near speed of light through an 844 metres long circle.
The subjected tissue is in a cryogenic condition that protects it from x-ray damaging. When the X-ray passes through, it captures subtle changes in passing tissues and this helps it to create the imaging.
The resulting images were interpreted by AI to distinguish nerves from tissues. In the study, scientists X-rayed millimetre size neural tissues and then built 3D images of them. Their resolutions were around 87 nanometres which could clearly display individual neurons and their pathways. All these were achieved within days where current procedures in use may take up to months or even years.
According to Wei-Chung Allen Lee, co-corresponding author in the study, “We think this is going to open new avenues for understanding the brain, both in how it’s organized and the circuitry that underlies its function. This type of knowledge can give us foundational insights into neurological disorders, diseases that affect the structure of the brain and much more.”